Welcome to the Bauer Lab
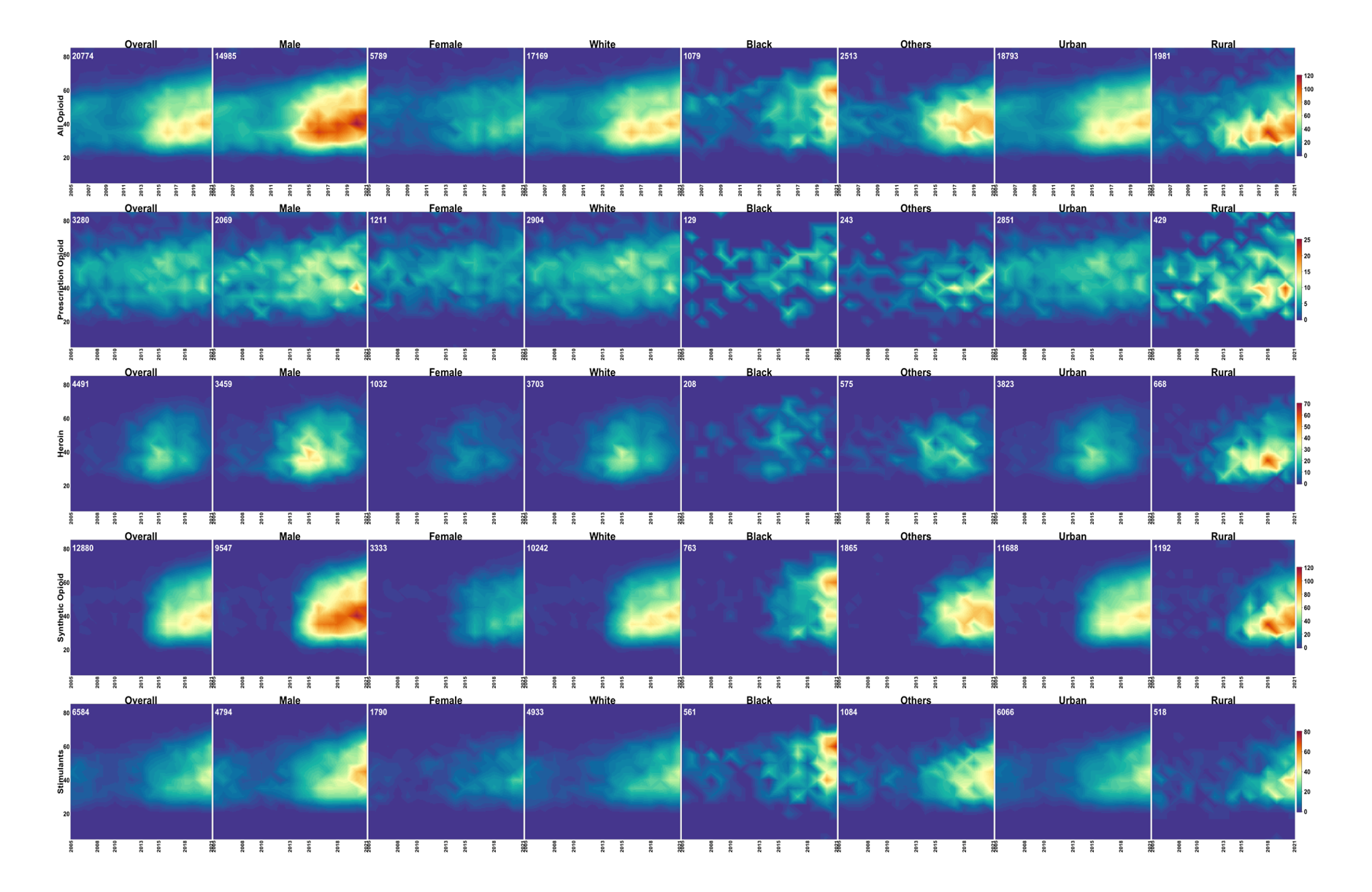
Dynamic temporal trends
Investigate the geospatial pattern of ALAN in US counties from 2012 to 2019.
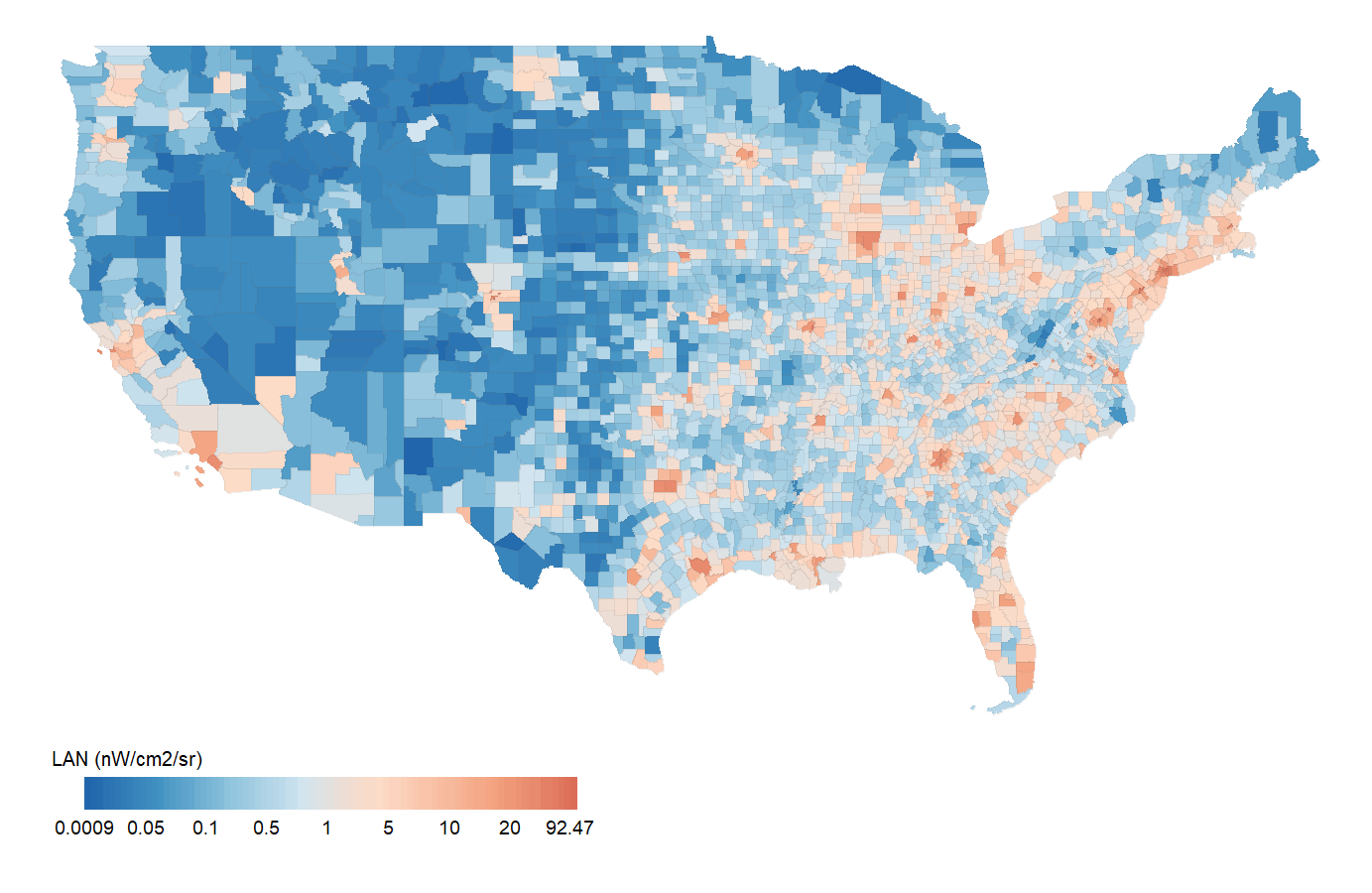
Artificial Light at Night
Investigate the geospatial pattern of ALAN in US counties from 2012 to 2019.